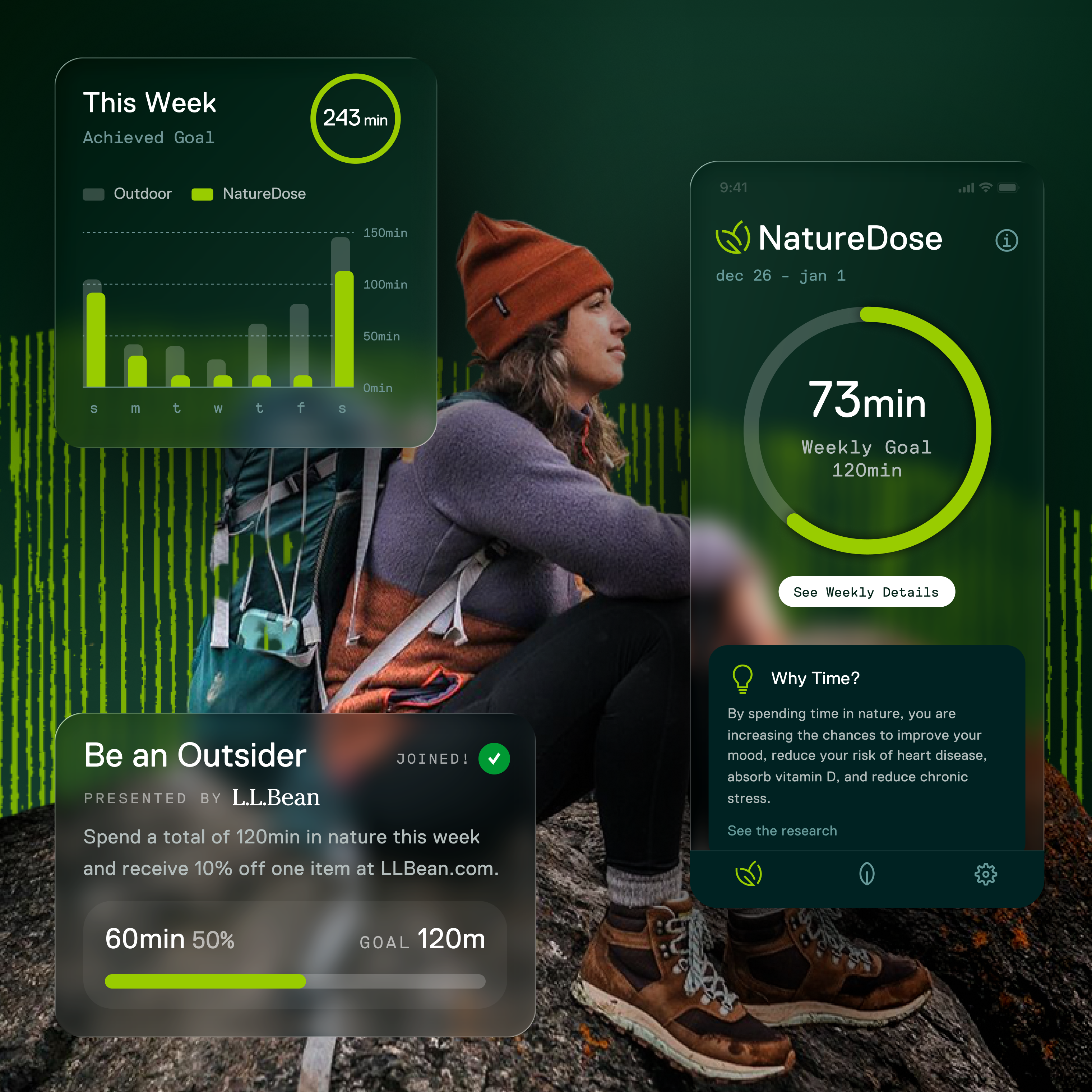
Visualization Community Designing a new way for scientists to see and share genomic discoveries.
Overview
Genomic visualizations at St. Jude were stuck — locked on local machines or flattened into static journal figures that lost all their interactivity the moment they were published. Researchers who wanted to revisit or share them had to start from scratch.
I led design for the Visualization Community, a platform on St. Jude Cloud that lets scientists publish, explore, and share living visualizations — without needing to learn how to write or run code specifically for ProteinPaint. What used to take weeks, now takes minutes. Just plug in your data's URL and visualize it with ProteinPaint.
Published visualizations produced across research projects
Visualizations featured in Nature, Cancer Discovery, and peer-reviewed journals
Research teams connected across institutions
Views for the leading publication visualization
Views for the AGI Community piece
Consecutive nominations for the Visualization is Beautiful Award
Challenges
Pediatric cancer research generates enormous, complex datasets. The insights are there — but making them visible, shareable, and useful to collaborators across the world was a different problem entirely.
Visualizations were flattened into static journal figures and lost their interactivity the moment they were published. Updating them required technical help. Sharing them meant starting over. The science was moving fast; the tools weren’t keeping up.
Contributions
St. Jude Cloud — already dedicated to open data — was the right foundation. The goal was a platform where researchers could publish, browse, and interact with visualizations without any technical setup. I designed the full product experience: identity, homepage, editor, track library, sample data views, and community pages.
Visual Identity
I designed the Visualization Community’s brand identity, including the logo and visual system that carries through the platform.
 Visualization Community logo
Visualization Community logo
 3D Genome Consortium logo — a collaboration between St. Jude and The Broad Institute at MIT
3D Genome Consortium logo — a collaboration between St. Jude and The Broad Institute at MIT
Platform Design
I designed the end-to-end platform experience — from the homepage and visualization explorer to the editor, sample data views, and community pages.
 Visualization editor — track configuration view
Visualization editor — track configuration view
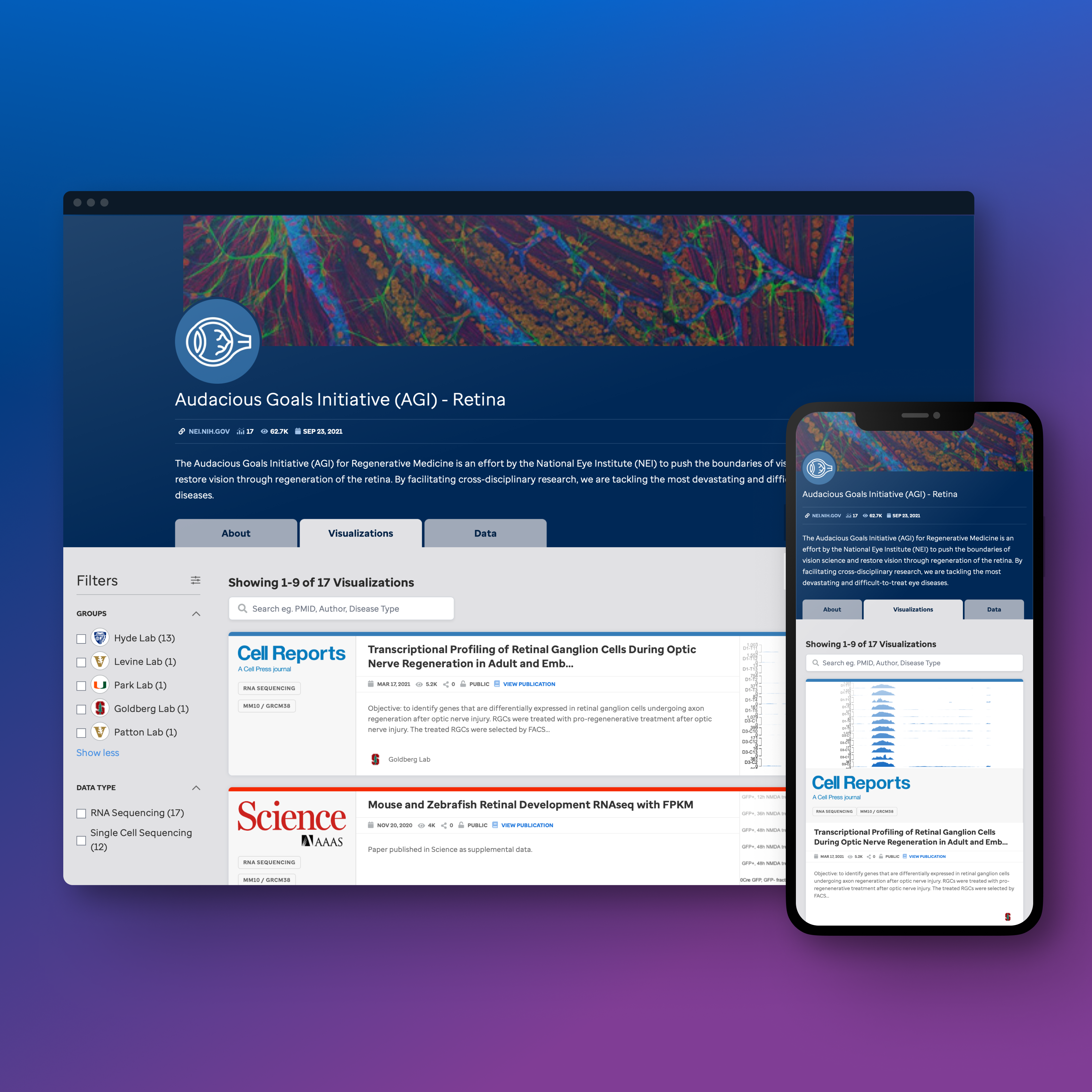 Communities view
Communities view
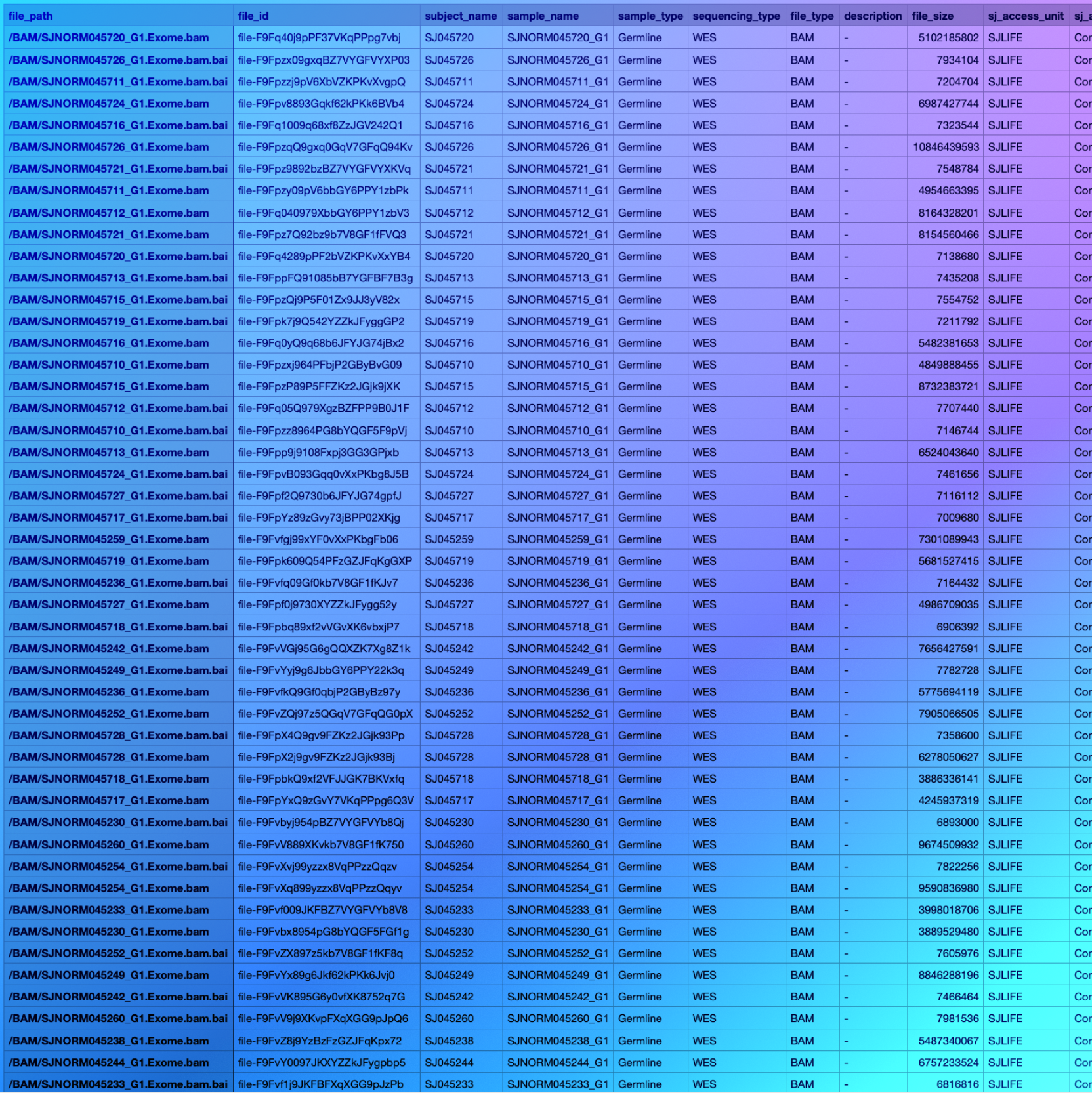 Sample data browser
Sample data browser
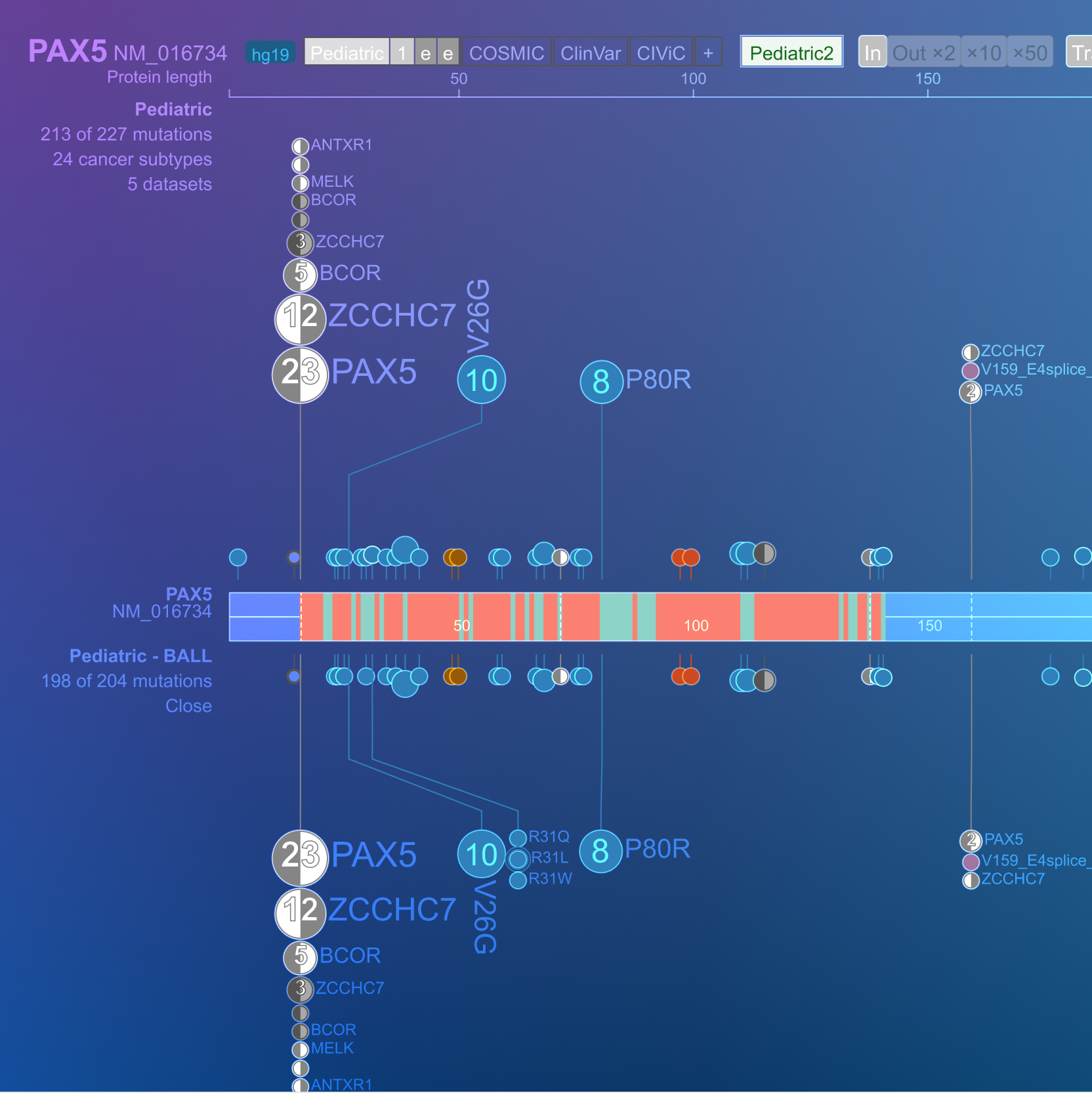 ProteinPaint visualization
ProteinPaint visualization
 Track component library
Track component library
"Anyone who goes into the visualization applications can look for the underlying data, analyze it, and publish using it if they find something novel. It's really a resource for the entire academic community."
Design
The platform supports a wide range of visualization types — from gene expression and mutation plots to genome-wide tracks. Each track type needed to be discoverable, well-labeled, and consistent in how it presented data.
I designed the track type library and the UI patterns for browsing, configuring, and publishing visualizations within the editor.
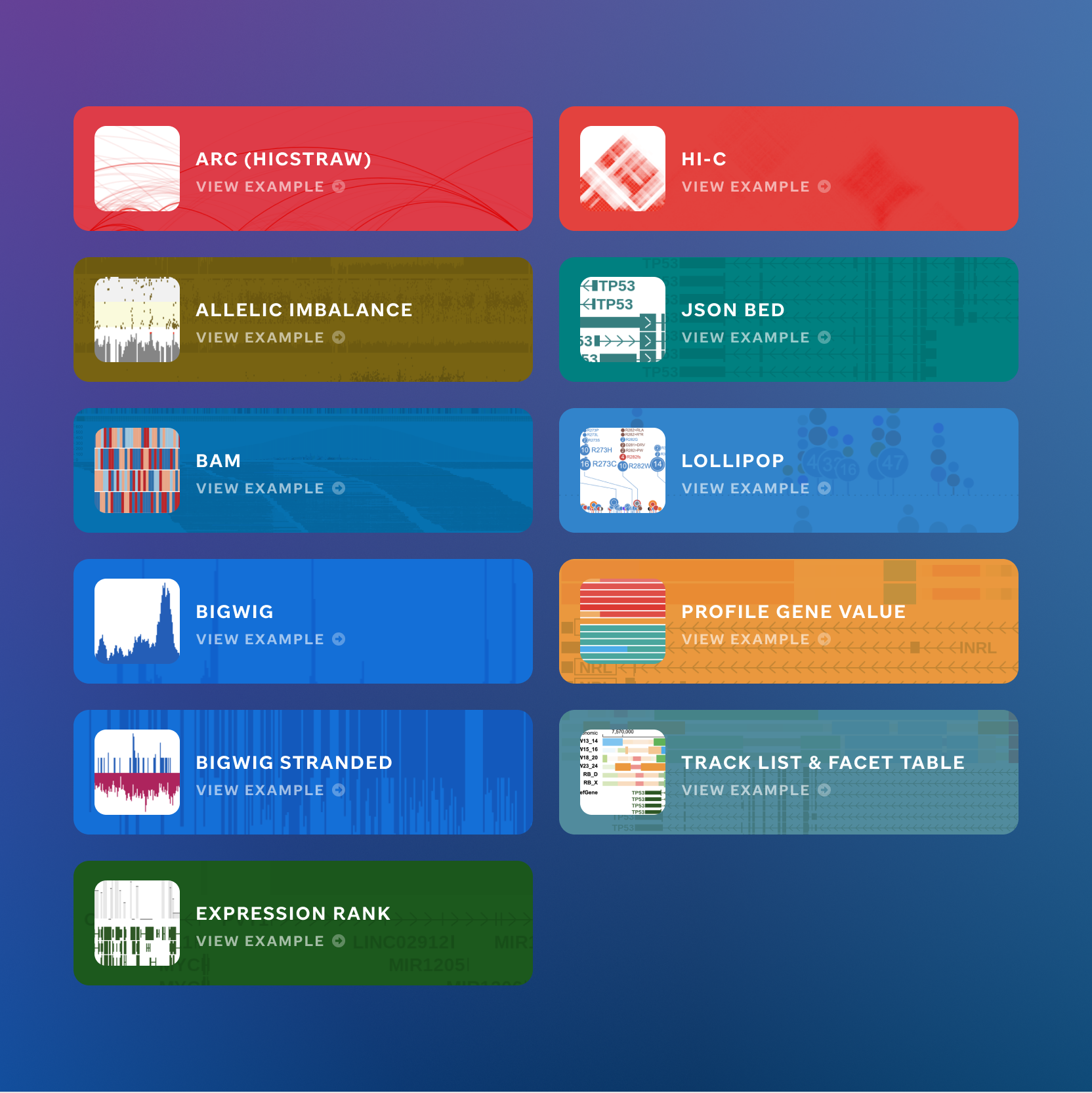 Visualization track type overview
Visualization track type overview
Code Editor
Each track type has a dedicated code editor view — letting researchers configure parameters and see live output as they build their visualization.
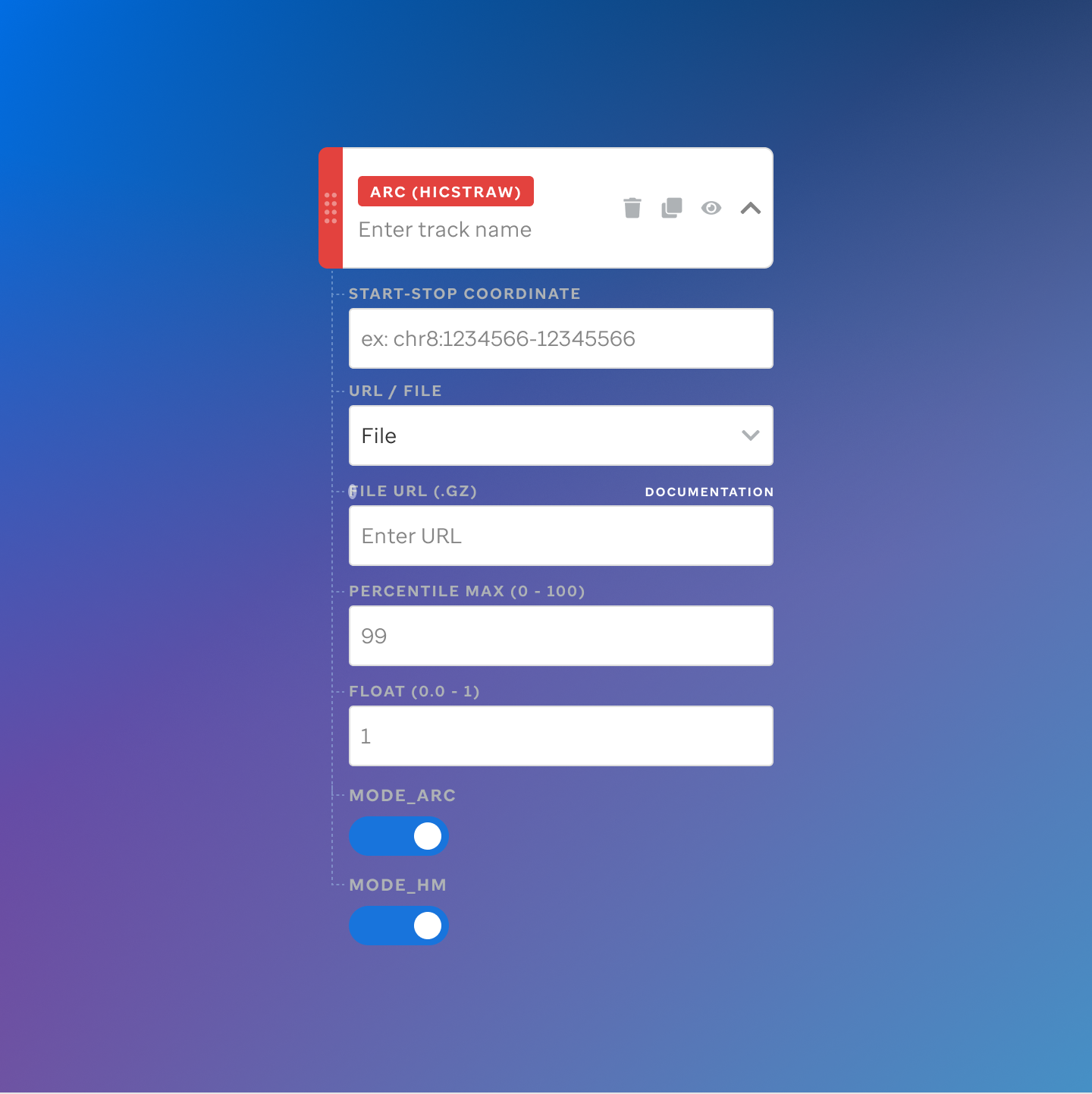
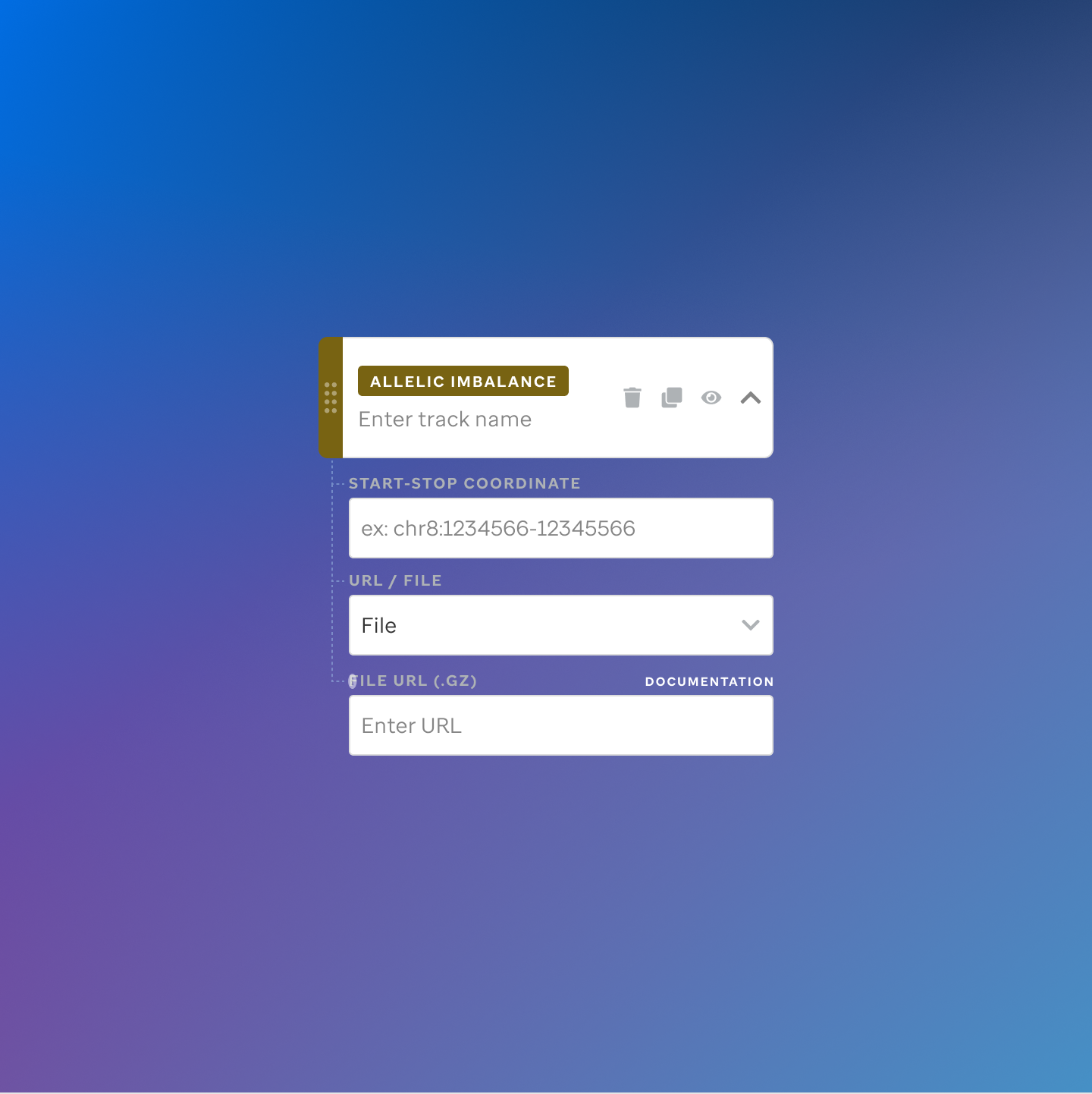
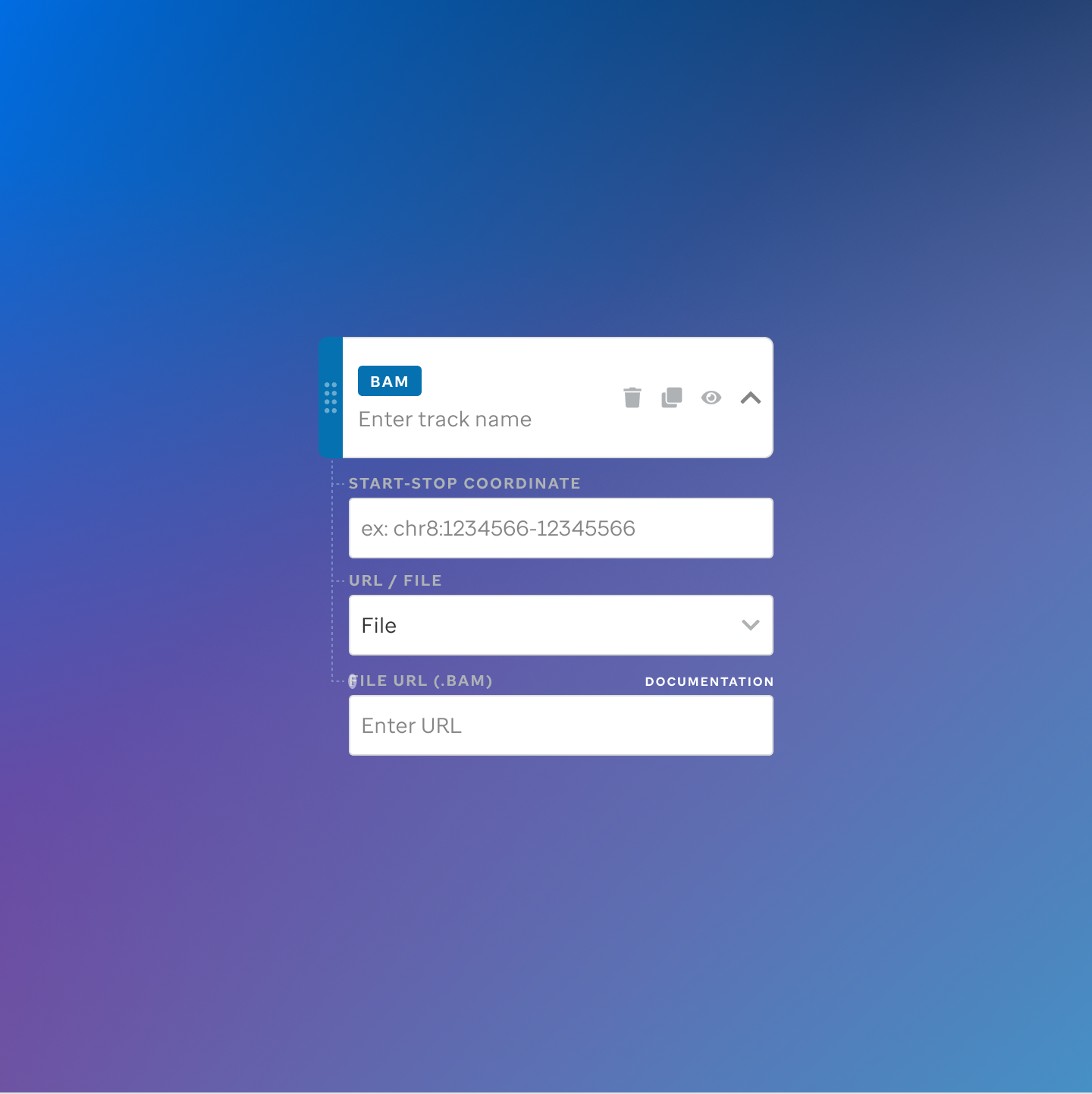
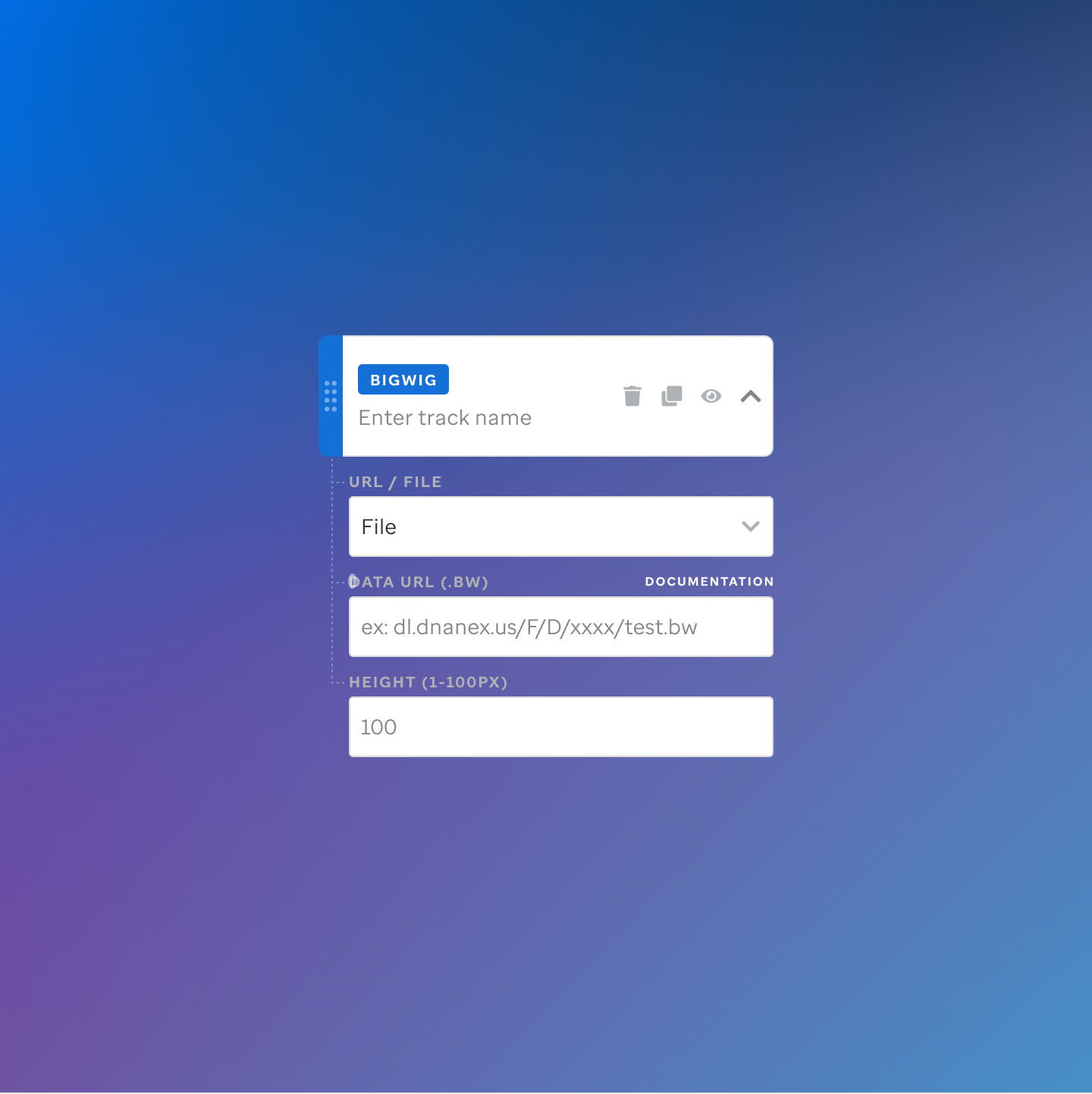
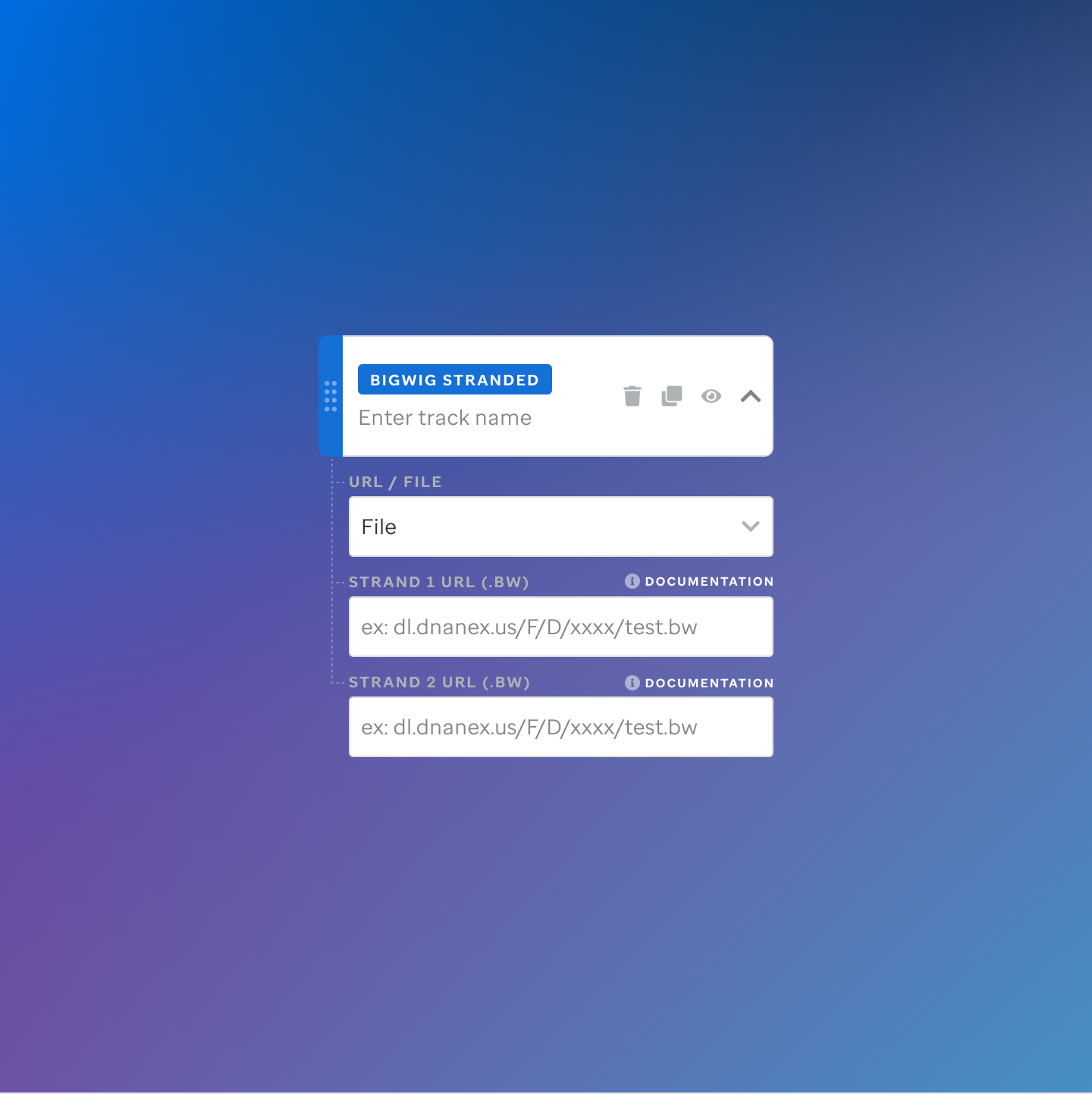
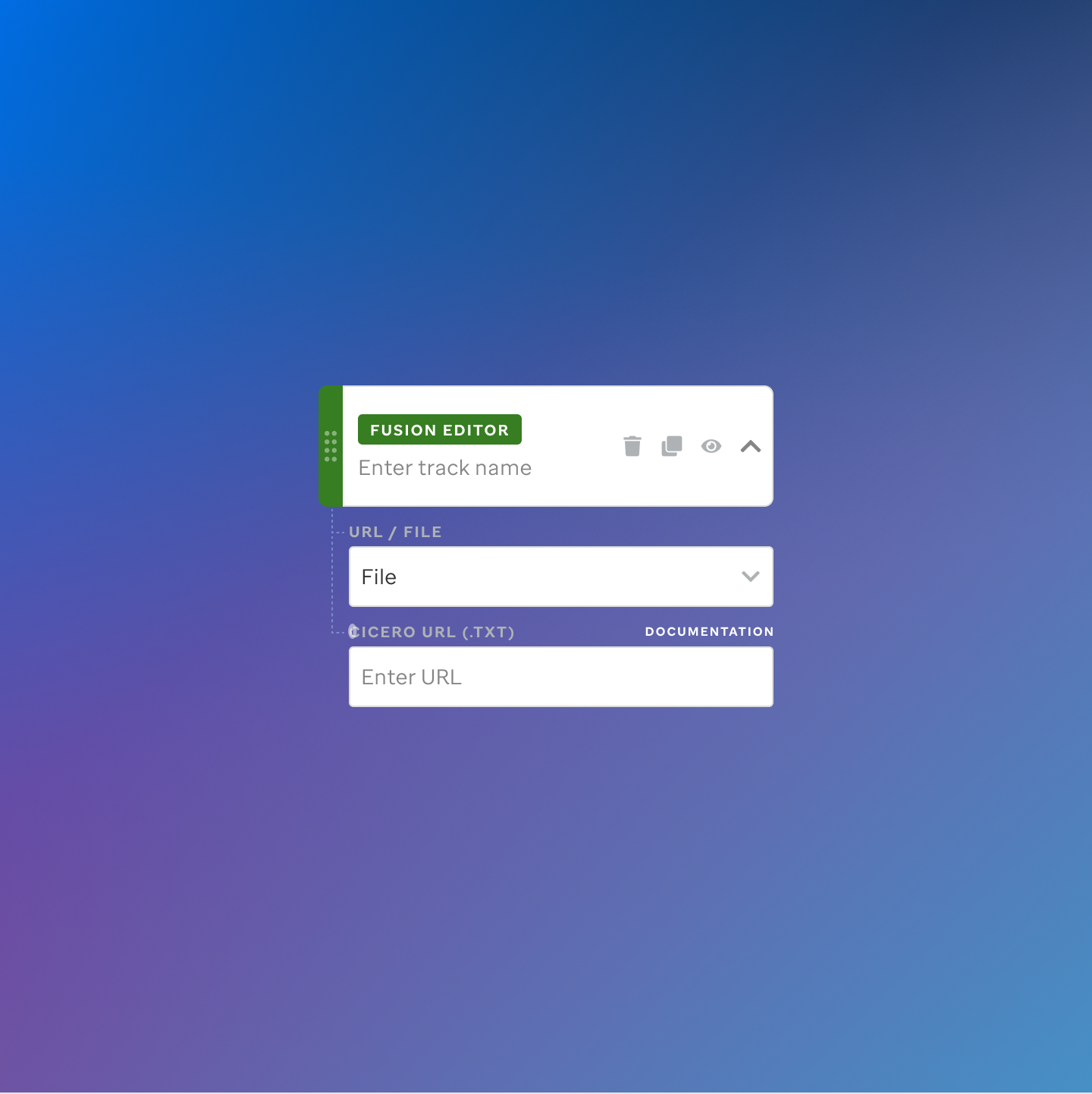
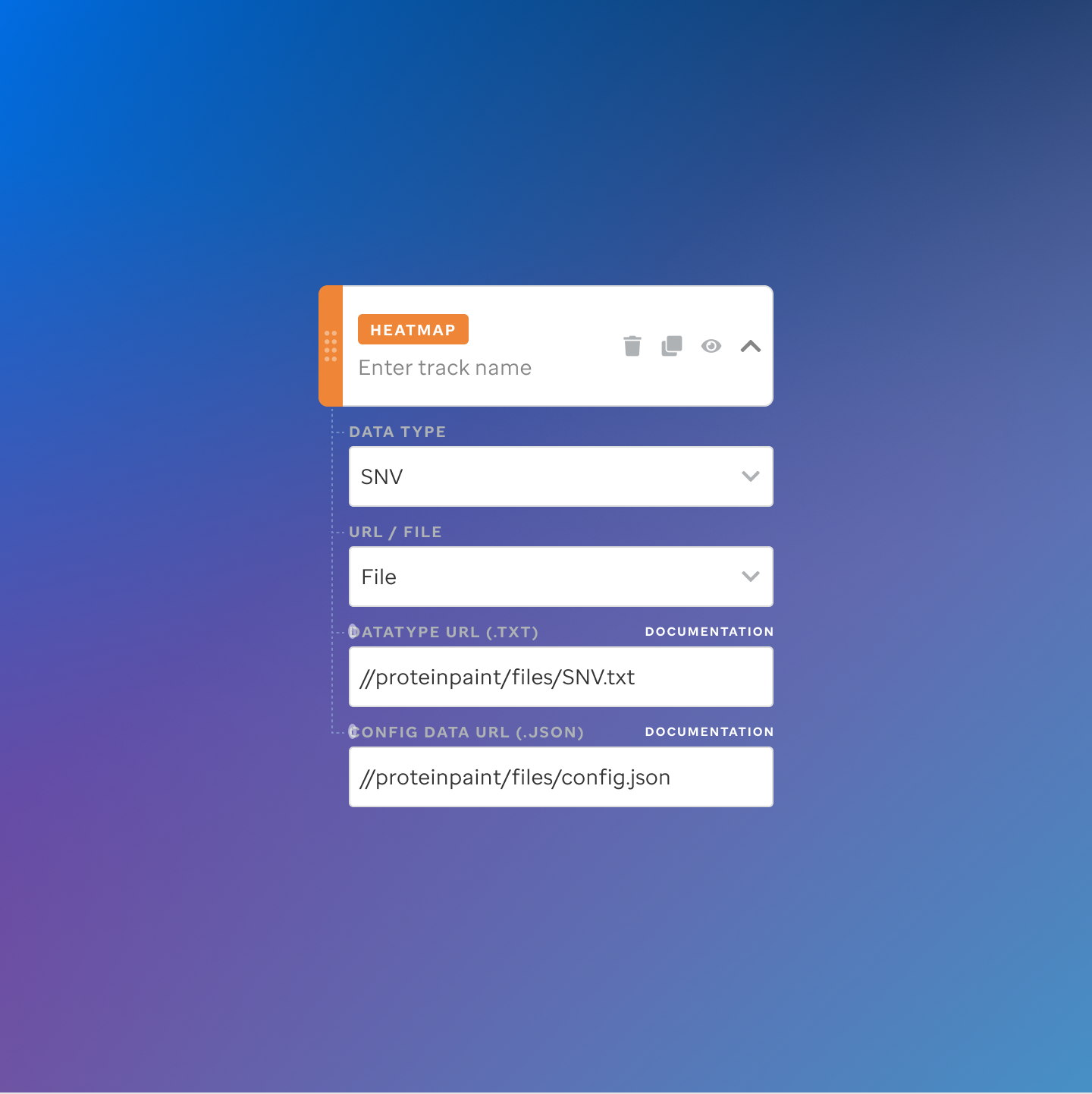
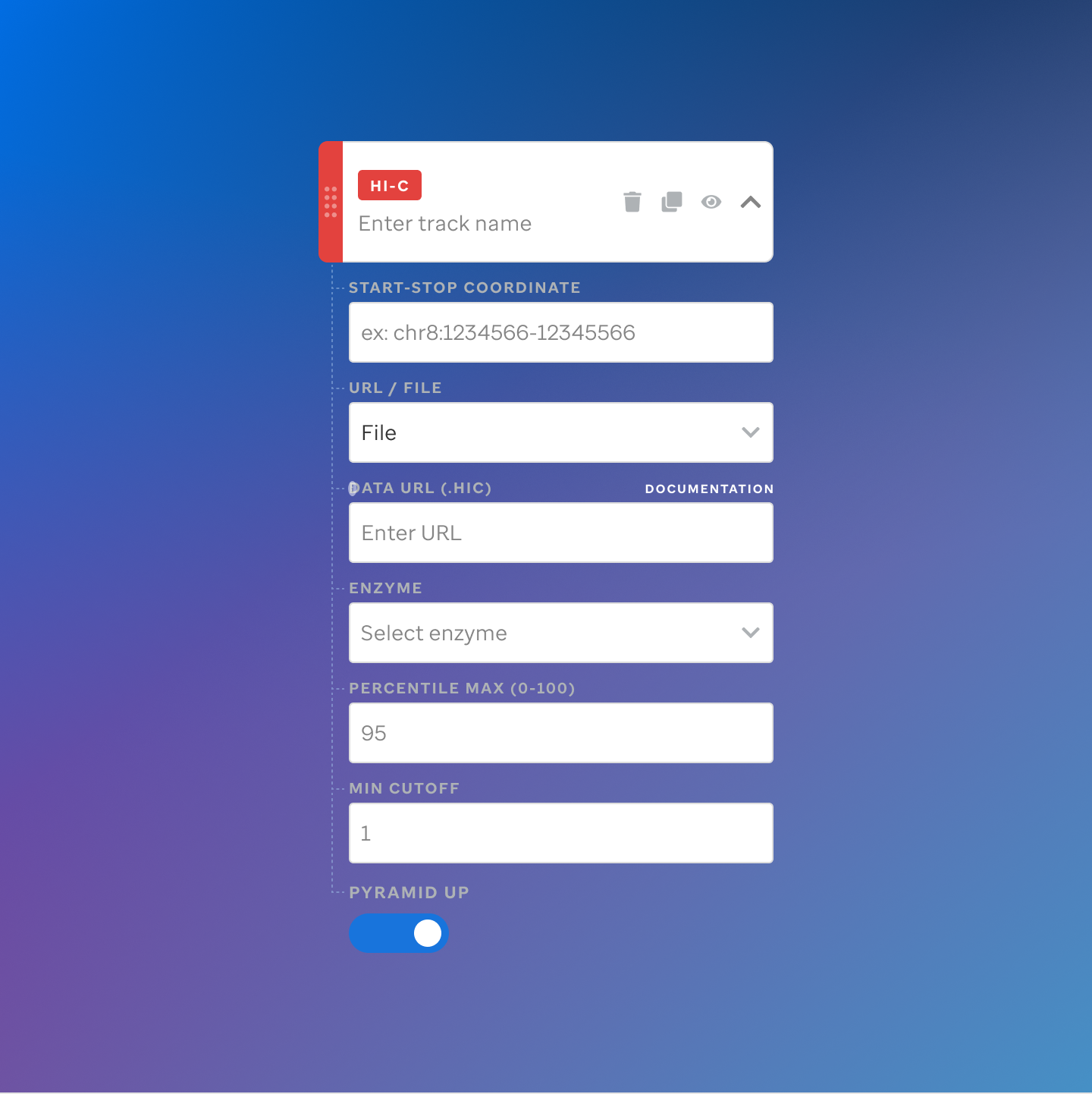
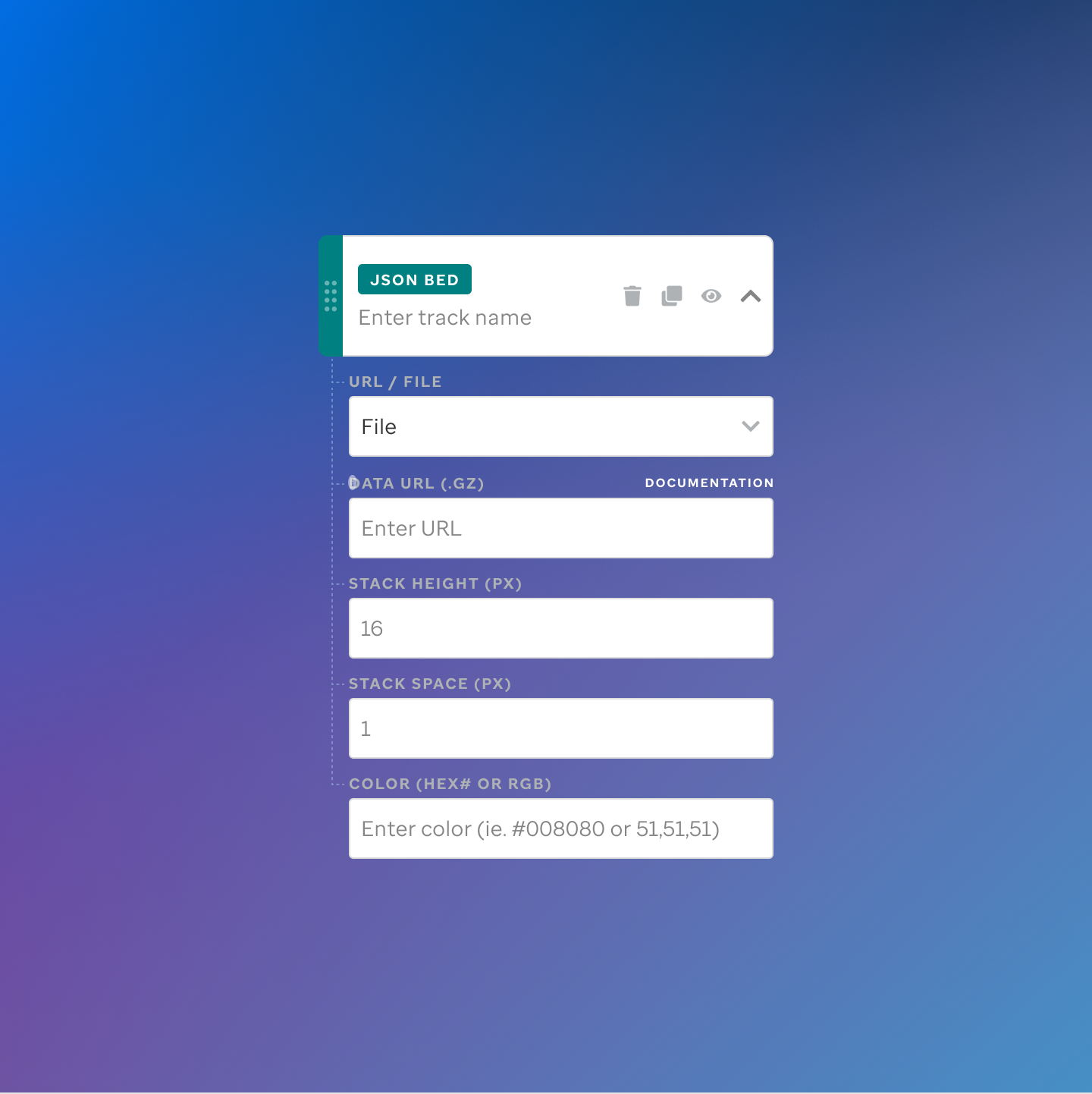
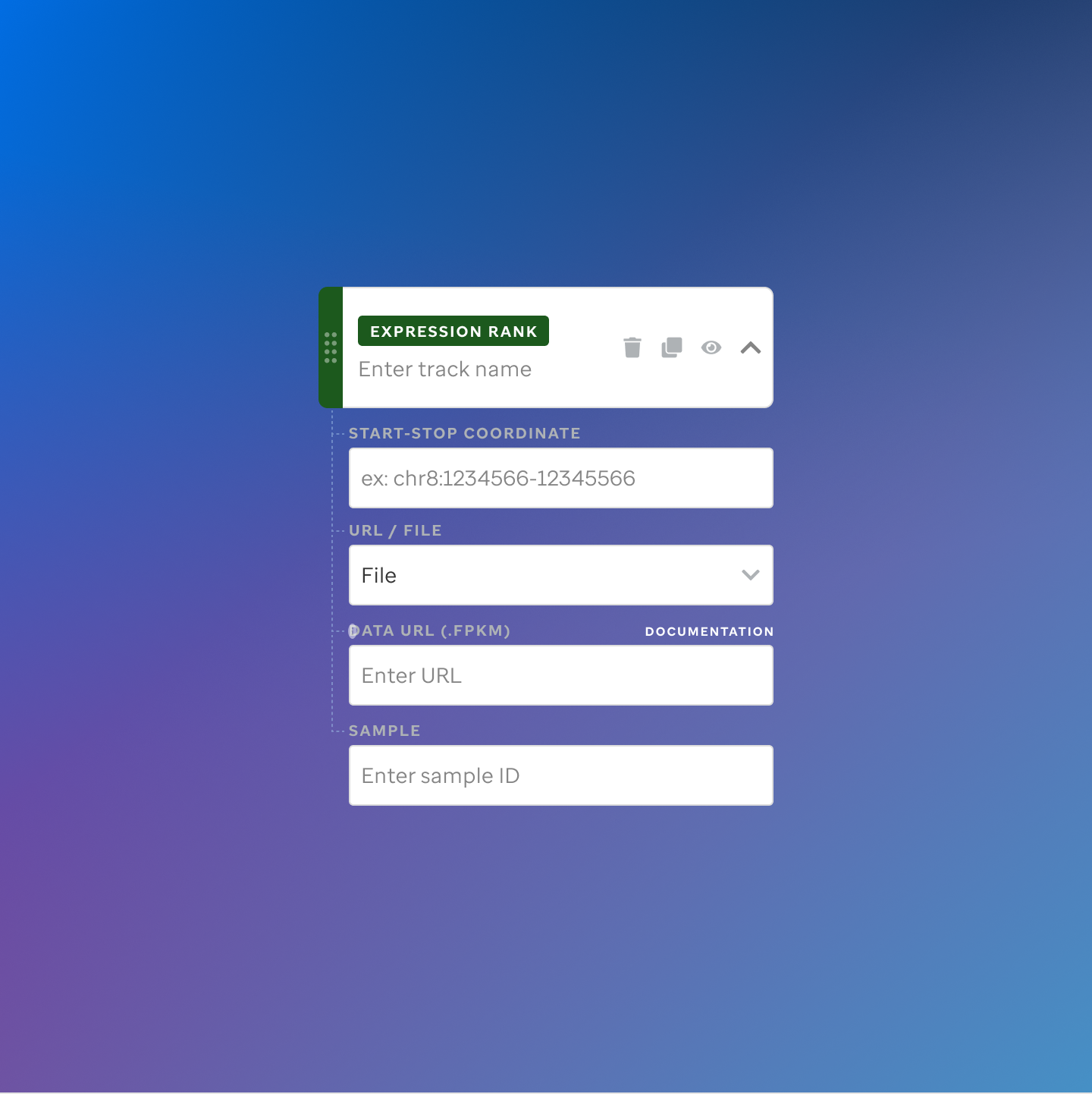
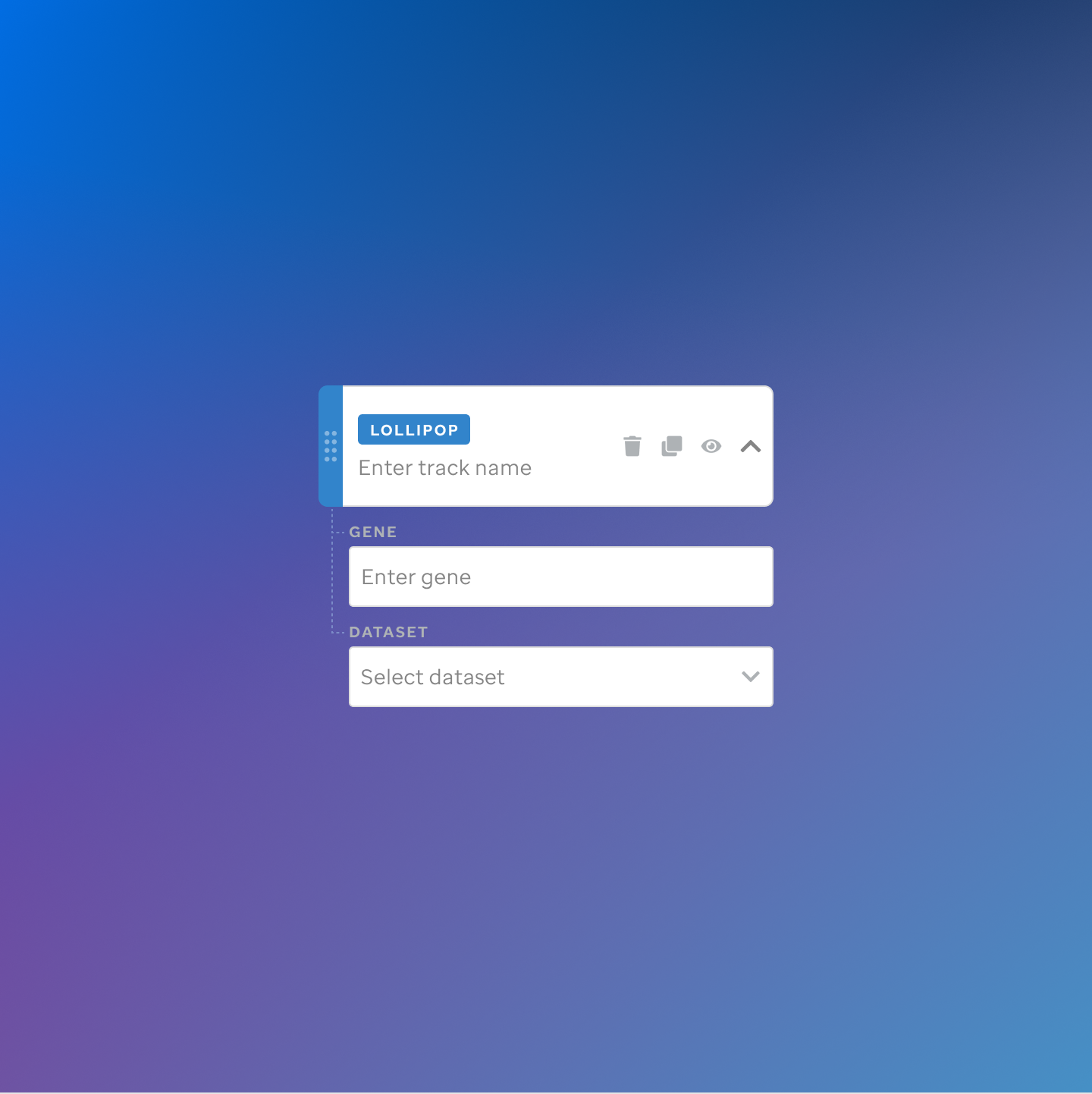
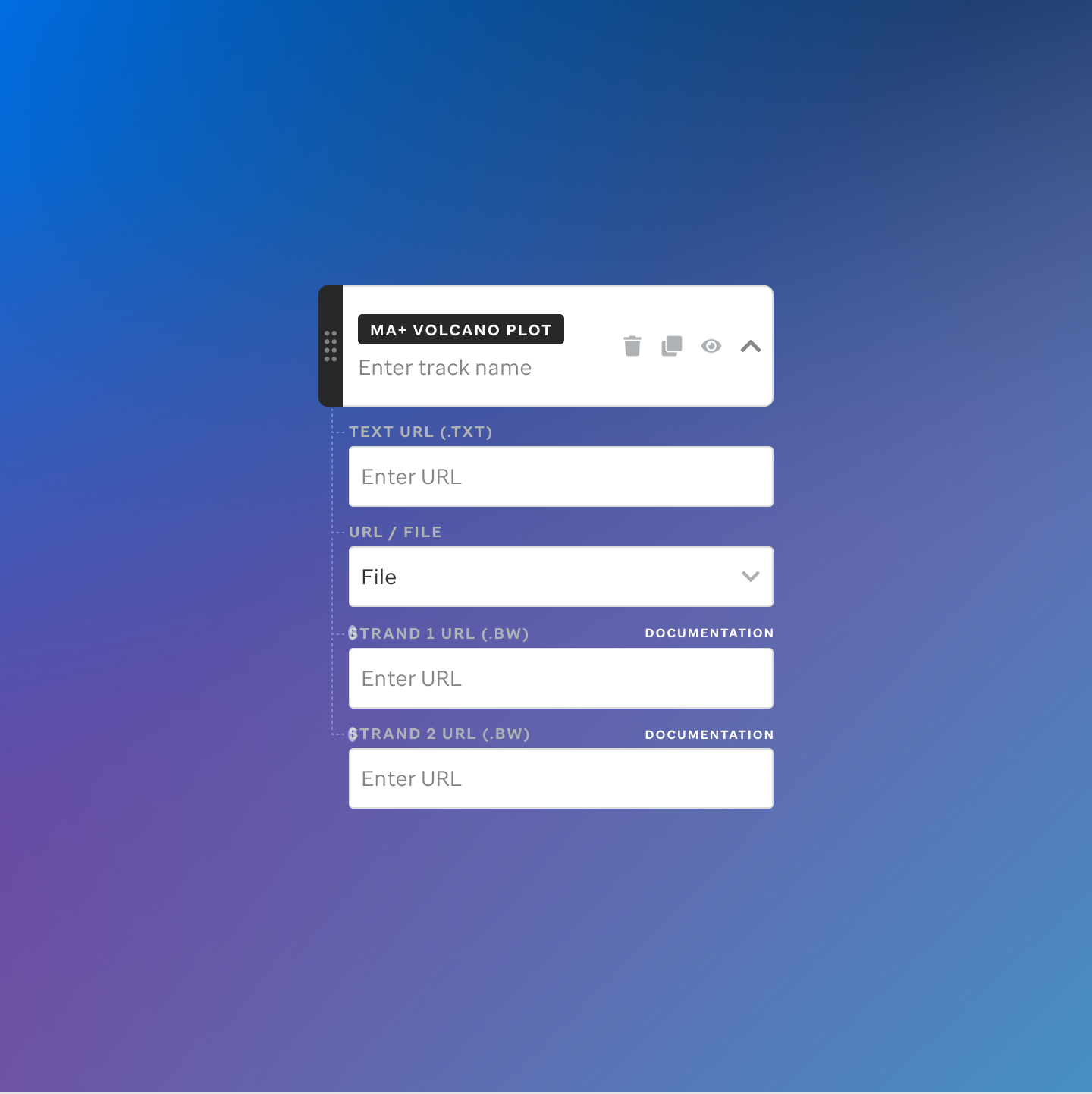
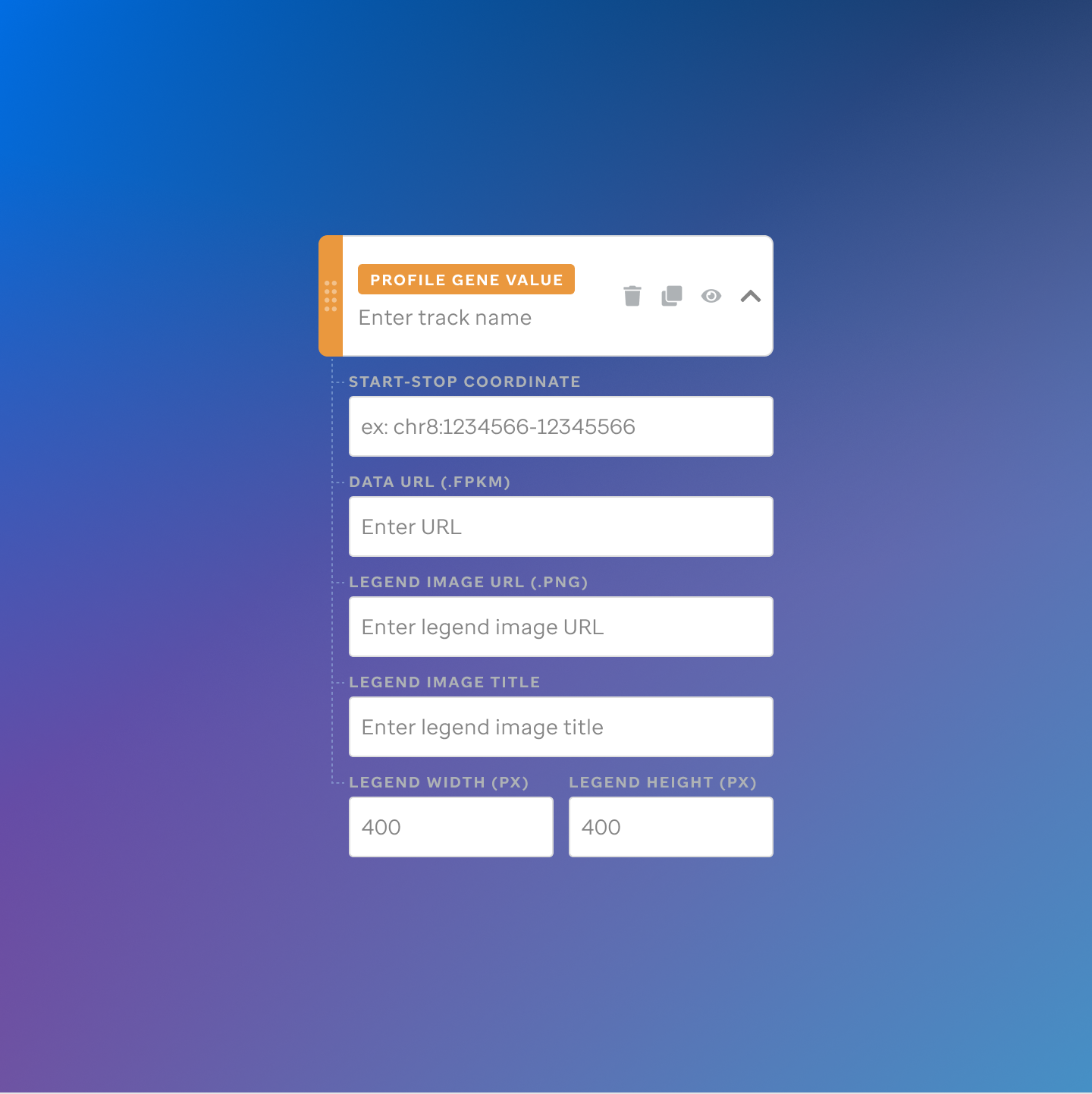
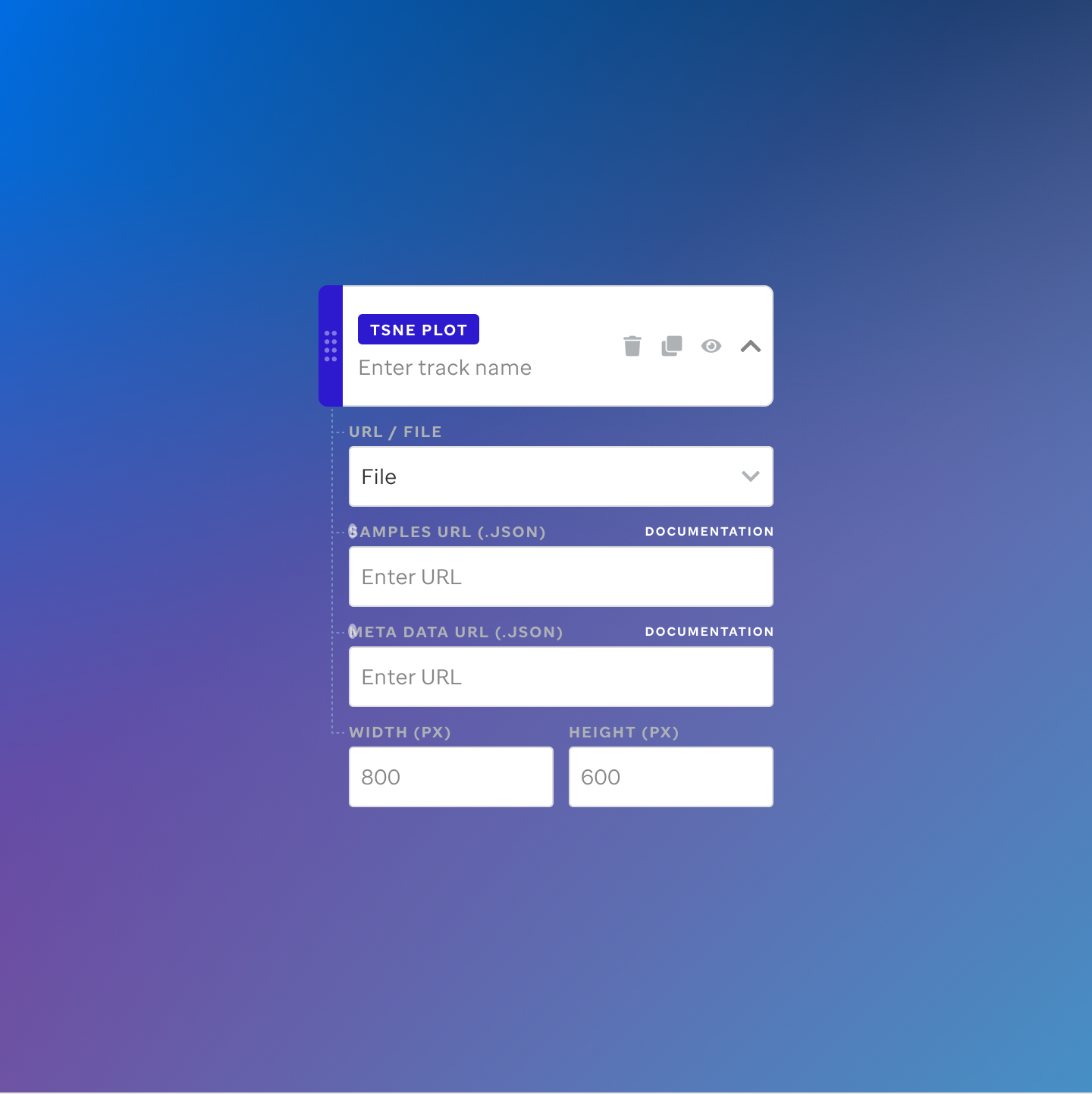
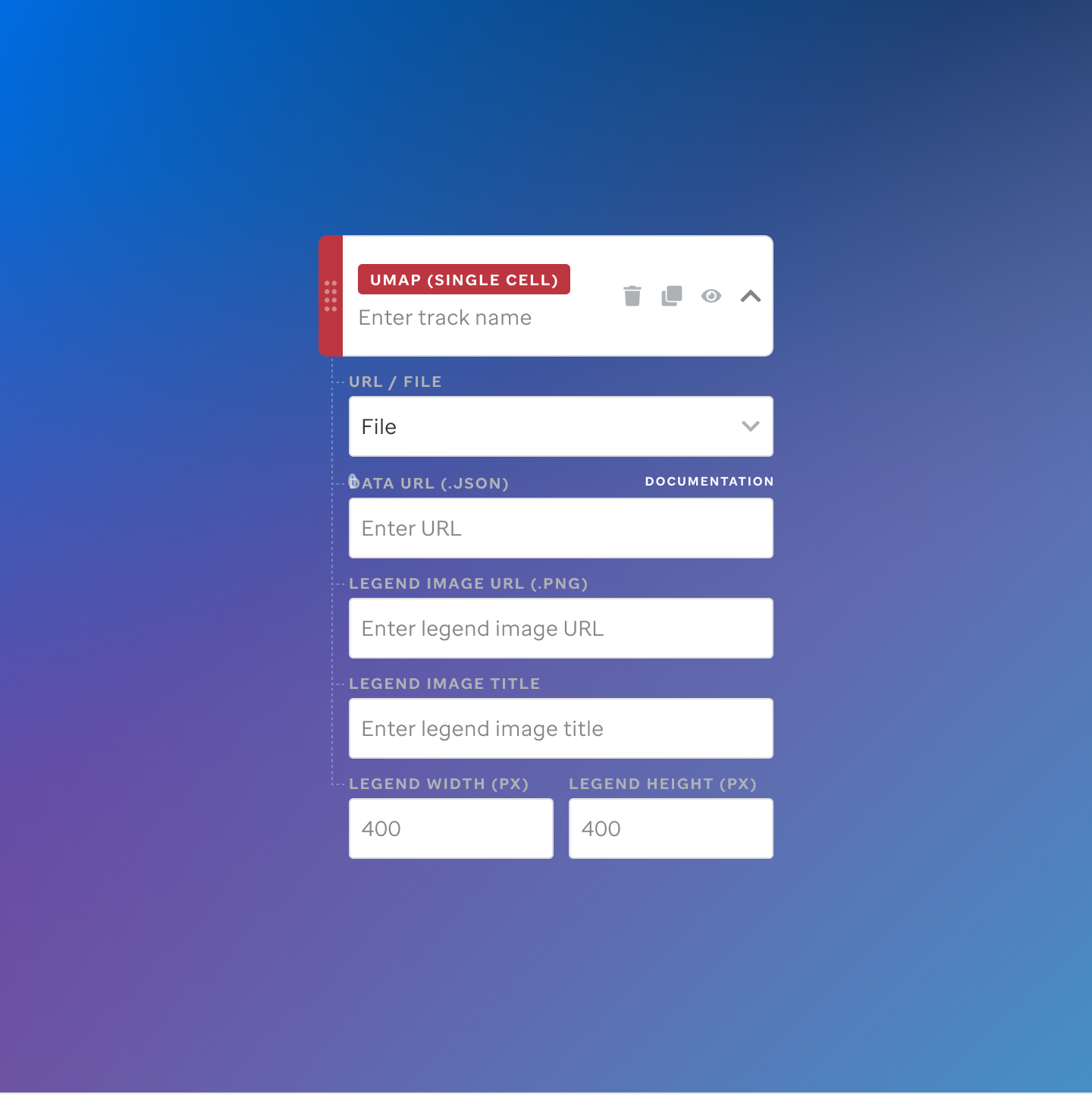
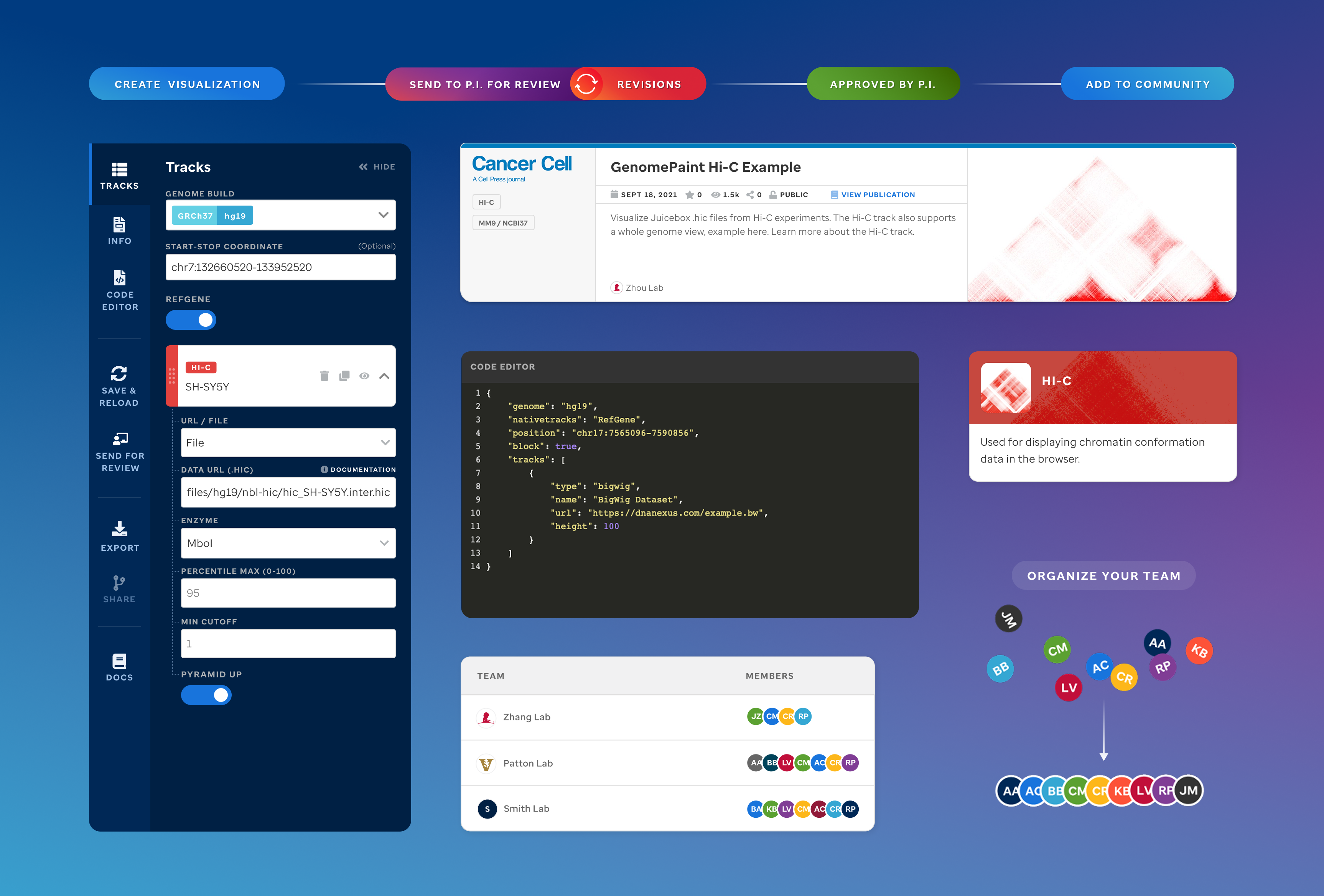 St. Jude Cloud — Visualization Community
St. Jude Cloud — Visualization Community
Impact
Researchers move faster.
They can build and update visualizations without waiting for technical help.
Access is reliable.
Data and visualizations now live in one stable, public home.
Insights run deeper.
Scientists can explore datasets in real time instead of relying on static images.
Impact extends further.
Teams around the world can now build on each other’s work.
 AACR conference poster presenting the Visualization Community platform
AACR conference poster presenting the Visualization Community platform
Reflection
Designing for researchers taught me that interface complexity has a real cost. When the tool is confusing, scientists ask for help instead of exploring. When it’s clear, they go further on their own.
The most meaningful moment on this project wasn’t a launch metric — it was watching a researcher configure and publish a visualization in a few minutes that would have taken a week before. That’s what the design was for.